Rows: 8,000
Columns: 10
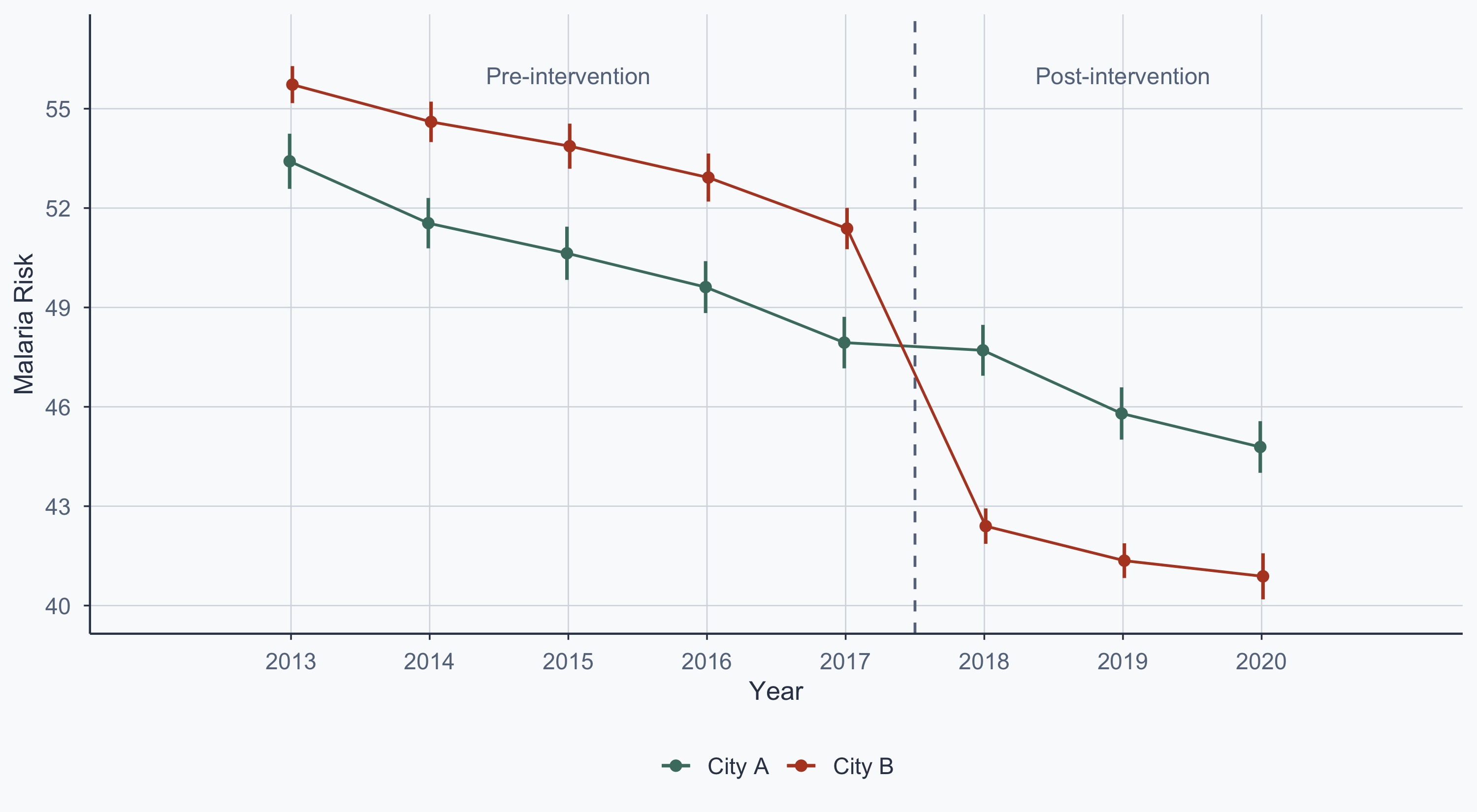
$ year <int> 2013, 2014, 2015, 2016, 2017, 2018, 2019, 2020, 2013, 201…
$ income <dbl> 1284.551, 1285.832, 1285.997, 1285.157, 1286.313, 1287.24…
$ age <int> 33, 34, 35, 36, 37, 38, 39, 40, 51, 52, 53, 54, 55, 56, 5…
$ sex <chr> "Woman", "Woman", "Woman", "Woman", "Woman", "Woman", "Wo…
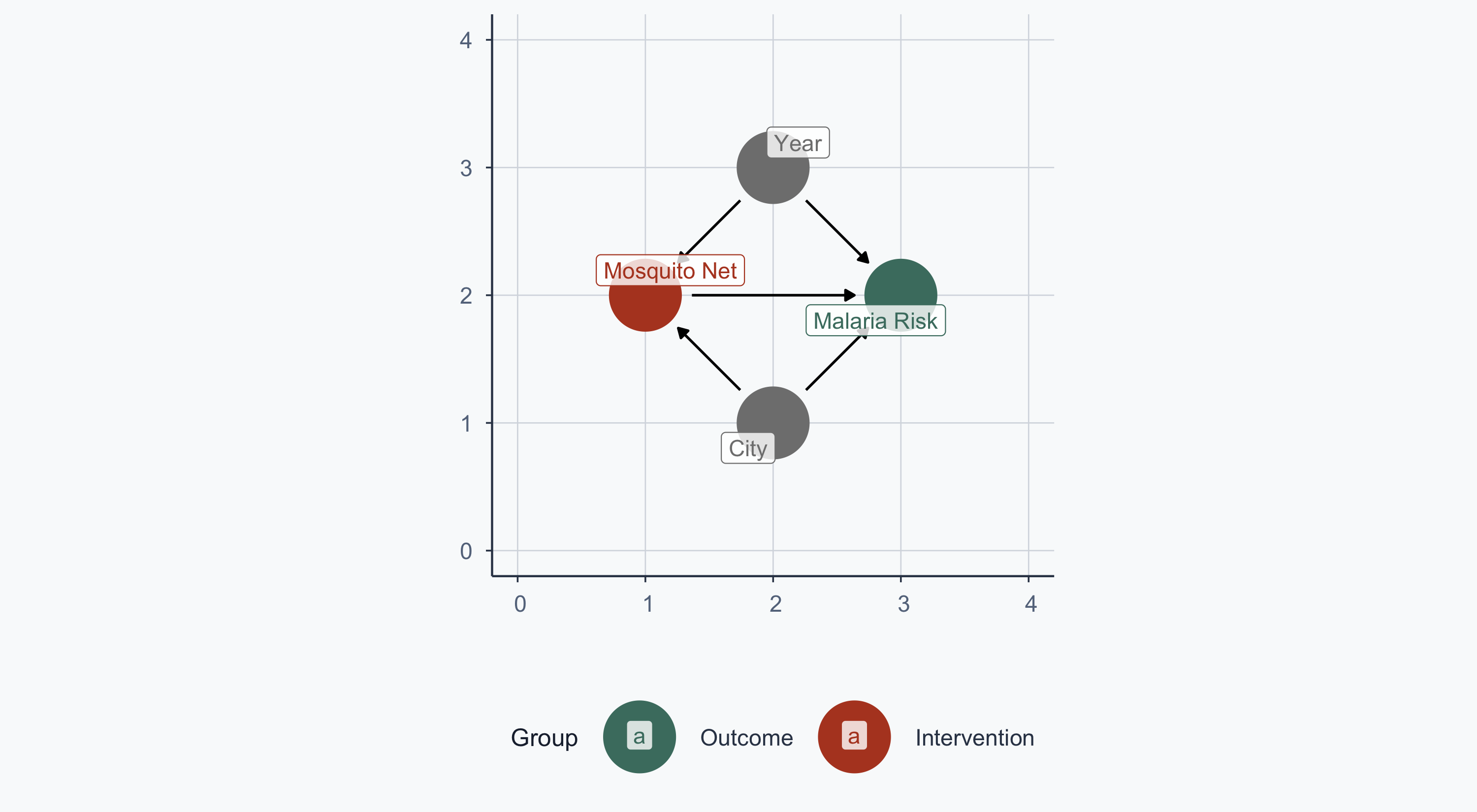
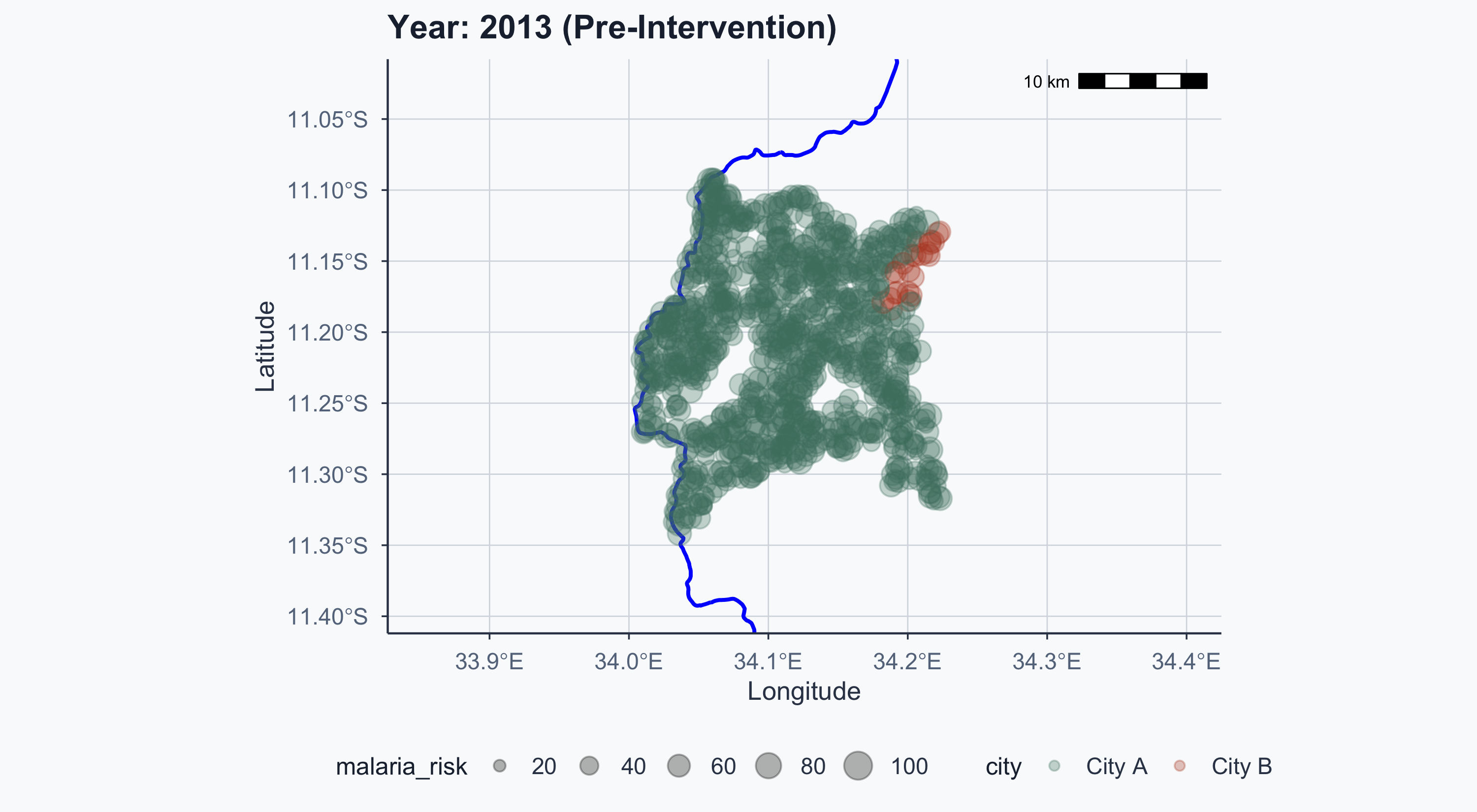
$ malaria_risk <dbl> 36.26529, 35.10382, 73.17664, 28.98390, 51.18721, 51.9071…
$ id <int> 1, 1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 2, 3, 3, 3, …
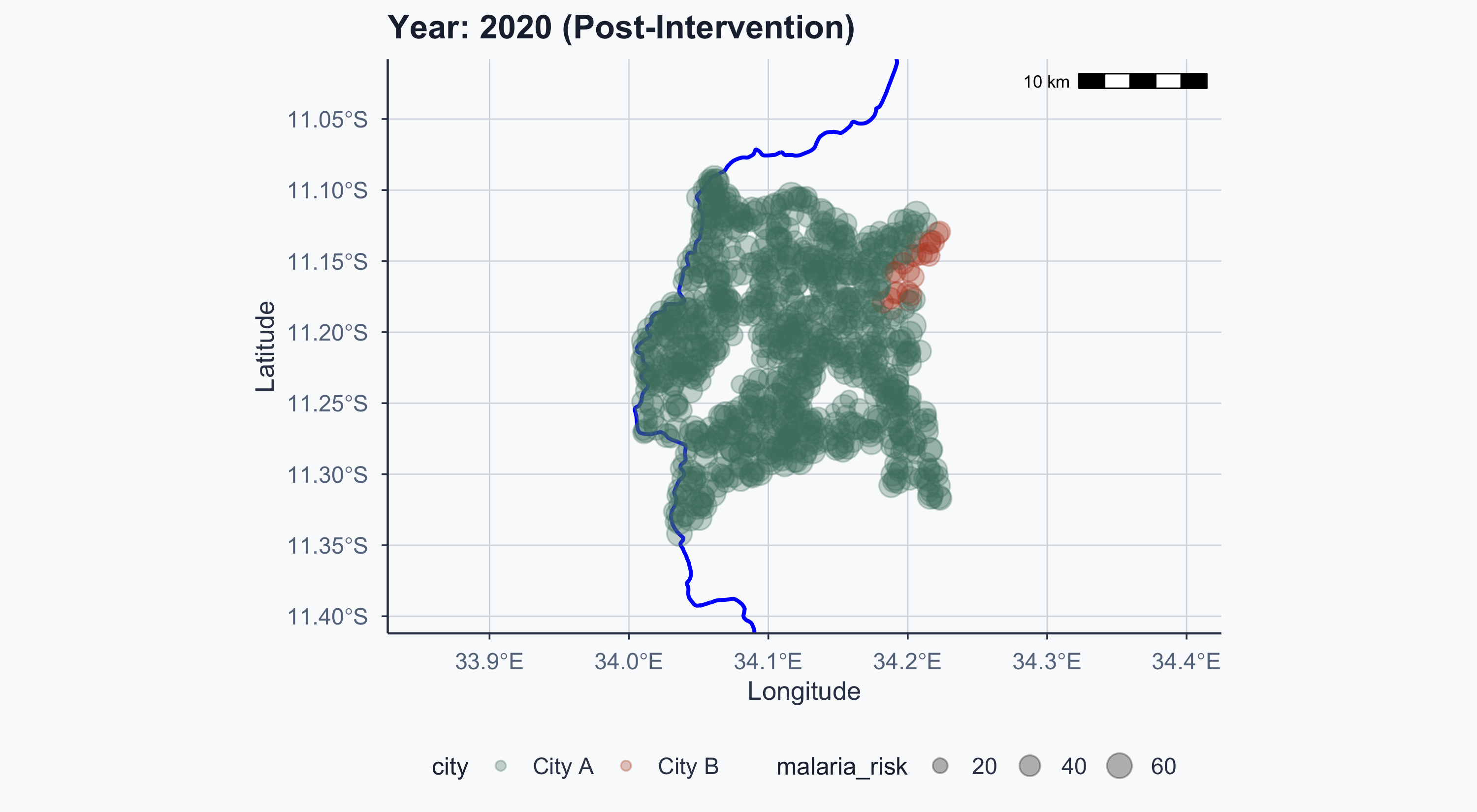
$ lat <dbl> -11.27476, -11.27476, -11.27476, -11.27476, -11.27476, -1…
$ lon <dbl> 34.03006, 34.03006, 34.03006, 34.03006, 34.03006, 34.0300…
$ after <int> 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, …
$ city <chr> "City A", "City A", "City A", "City A", "City A", "City A…